I spent the last two weeks teaching frequentist and Bayesian statistics at the
European Summer School in Logic, Language, and Information (ESSLLI) in Barcelona, at the beautiful and centrally located Pompeu Fabra University. The course web page for the first week is
here, and the web page for the second course is
here.
All materials for the first week are available on github, see
here.
The frequentist course went well, but the Bayesian course was a bit unsatisfactory; perhaps my greater experience in teaching the frequentist stuff played a role (I have only taught Bayes for three years). I've been writing and rewriting my slides and notes for frequentist methods since 2002, and it is only now that I can present the basic ideas in five 90 minute lectures; with Bayes, the presentation is more involved and I need to plan more carefully, interspersing on-the-spot exercises to solidify ideas. I will comment on the Bayesian Data Analysis course in a subsequent post.
The first week (five 90 minute lectures) covered the basic concepts in frequentist methods. The audience was amazing; I wish I always had students like these in my classes. They were attentive, and anticipated each subsequent development. This was the typical ESSLLI crowd, and this is why teaching at ESSLLI is so satisfying. There were also several senior scientists in the class, so hopefully they will go back and correct the misunderstandings among their students about what all this Null Hypothesis Significance Testing stuff gives you (short answer: it answers *a* question very well, but it's the wrong question, nothing that is relevant to your research question).
I won't try to summarize my course, because the web page is online and you can also do exercises on datacamp to check your understanding of statistics (see
here). You get immediate feedback on your attempts.
Stepping away from the technical details, I tried to make three broad points:
First, I spent a lot of time trying to clarify what a p-value is and isn't, focusing particularly on the issue of Type S and Type M errors, which Gelman and Carlin have discussed in
their excellent paper.
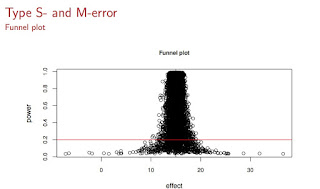
Here is the way that I visualized the problems of Type S and Type M errors:
What we see here is repeated samples from a Normal distribution with true mean 15 and a typical standard deviation seen in psycholinguistic studies (see slide 42 of my slides for lecture2). The horizontal red line marks the 20% power line; most psycholinguistic studies fall below that line in terms of power. The dramatic consequence of this low power is the hugely exaggerated effects (which tend to get published in major journals because they also have low p-values) and the remarkable proportion of cases where the sample mean is on the wrong side of the true value 15. So, you are roughly equally likely to get a significant effect with a sample mean smaller to much smaller than the true mean, or larger and much larger than the true mean. With low power, regardless of whether you get a significant result or not, if power is low, and it is in most studies I see in journals, you are just farting in a puddle.
It is worth repeating this: once one considers Type S and Type M errors, even statistically significant results become irrelevant, if power is low. It seems like these ideas are forever going to be beyond the comprehension of researchers in linguistics and psychology, who are trained to make binary decisions based on p-values, weirdly accepting the null if p is greater than 0.05 and, just as weirdly, accepting their
favored alternative if p is less than 0.05. The p-value is a truly interesting animal; it seems tha
t a recent survey of some 400 Spanish psychologists found that, despite their being active in the field for quite a few years on average, they had close to zero understanding of what a p-value gives you. Editors of top journals in psychology routinely favor lower p-values, because they mistakenly think this makes "the result" more convincing; "the result" is the favored alternative. So even seasoned psychologists (and I won't even get started with linguists, because we are much, much worse), with decades of experience behind them, often have no idea what the p-value actually tells you.
A remarkable misunderstanding regarding p-values is the claim that it tells you whether the effect was "by chance". Here is an example from
Replication Index's blog:
"The Test of Insufficient Variance (TIVA) shows that the variance in z-scores is less than 1, but the probability of this event to occur by chance is 10%, Var(z) = .63, Chi-square (df = 11) = 17.43, p = .096."
Here is another example from a self-published manuscript by Daniel Ezra Johnson:
"If we perform a likelihood-ratio test, comparing the
model with gender to a null model with no predictors, we
get a p-value of 0.0035. This implies that it is very
unlikely that the observed gender difference is due to
chance."
One might think that the above examples are not peer-reviewed, and that peer review would catch such mistakes. But even people explaining p-values in publications are unable to understand that this is completely false. An example is Keith Johnson's textbook, Quantitative Methods in Linguistics, which repeatedly talks about "reliable effects" and effects which are and are not due to chance. It is no wonder that the poor psychologist/linguist thinks, ok, if the p-value is telling me the probability that the effect is due to chance, and if the p-value is low, then the effect is not due to chance and the effect must be true. The mistake here is that the p-value is telling you the probability of the result being "due to chance" conditional on the null hypothesis being true. It's better to explain the p-value as the probability of getting the statistic (e.g., t-value) or something more extreme,
under the assumption that the null hypothesis is true. People seem to drop the italicized part and this starts to propagate the misunderstanding for future generations. To repeat, the p-value is a conditional probability, but most people interpret it as an unconditional probability because they drop the phrase "under the null hypothesis" and truncate the statement to be about effects being due to chance.
Another bizarre thing I have repeatedly seen is misinterpreting the p-value as Type I error. Type I error is fixed at a particular value (0.05) before you run the experiment, and is the probability of your incorrectly rejecting the null when it's true, under repeated sampling. The p-value is what you get from your single experiment and is the conditional probability of your getting the statistic you got or something more extreme,
assuming that the null is true. Even this point is beyond comprehension for psychologists (and of course linguists).
Here is a bunch of psychologists explaining in an article why a p=0.000 should not be reported as an exact value:
"p = 0.000. Even though this statistical expression,
used in over 97,000 manuscripts according to Google Scholar,
makes regular cameo appearances in our computer printouts, we
should assiduously avoid inserting it in our Results sections. This
expression implies erroneously that there is a zero probability
that the investigators have committed a Type I error, that is, a
false rejection of a true null hypothesis (Streiner, 2007). That
conclusion is logically absurd, because unless one has examined
essentially the entire population, there is always some chance
of a Type I error, no matter how meager. Needless to say, the
expression “p < 0.000” is even worse, as the probability of
committing a Type I error cannot be less than zero. Authors
whose computer printouts yield significance levels of p = 0.000
should instead express these levels out to a large number of
decimal places, or at least indicate that the probability level is
below a given value, such as p < 0.01 or p < 0.001."
The p-value is the probability of committing a Type I error, eh? It is truly embarrassing that people who are teaching this stuff have distorted the meaning of the p-value so drastically and just keep propagating the error. I should mention though that this paper I am citing appeared in Frontiers, which I am beginning to question as a worthwhile publication venue. Who did the peer review on this paper and why did they not catch this basic mistake?
Even Fisher (
p. 16 of The Design of Experiments, Second Edition, 1937) didn't buy the p-value; he is advocating for replicability as the real decisive tool:
"It is usual and convenient for experimenters to take-5 per cent. as a standard level of significance, in the sense that they are prepared to ignore all results which fail to reach this standard, and, by this means, to eliminate from further discussion the greater part of the fluctuations which chance causes have introduced into their experimental results. No such selection can eliminate the whole of the possible effects of chance. coincidence, and if we accept this convenient convention, and agree that an event which would occur by chance only once in 70 trials is decidedly" significant," in the statistical sense, we thereby admit that no isolated experiment, however significant in itself, can suffice for the experimental demonstration of any natural phenomenon; for the "one chance in a million" will undoubtedly occur, with no less and no more than its appropriate frequency, however surprised we may be that it should occur to us. In order to assert that a natural phenomenon is experimentally demonstrable we need, not an isolated record, but a reliable method of procedure. In relation to the test of significance, we may say that a phenomenon is experimentally demonstrable when we know how to conduct an experiment which will rarely fail to give us a statistically significant result."
Second, I tried to clarify what a 95% confidence interval is and isn't. At least a couple of students had a hard time accepting that the 95% CI refers to the procedure and not that the true $\mu$ lies within one specific interval with probability 0.95, until I pointed out that $\mu$ is just a point value and doesn't have a probability distribution associated with it. Morey and Wagenmakers and Rouder et al have been shouting themselves hoarse about confidence intervals, and
how many people don't understand them, also see
this paper. Ironically, psychologists have responded to these complaints through various media, but even this response only showcases how psychologists have only a partial and misconstrued understanding of confidence intervals. I feel that part of the problem is that scientists hate to back off from a position they have taken, and so they tend to hunker down and defend defend defend their position. From the perspective of a statistician who understands both Bayes and frequentist positions, the conclusion would have to be that Morey et al are right, but for large sample sizes, the difference between a credible interval and a confidence interval (I mean the actual values that you get for the lower and upper bound) are negligible. You can see examples in our
recently ArXiv'd paper.
Third, I tried to explain that there is a cultural difference between statisticians on the one hand and (most) psychologists and almost all psychologist, linguists, etc. on the other. For the latter group (with the obvious exception of people using Bayesian methods for data analysis), the whole point of fitting a statistical model is to do a hypothesis test, i.e., to get a p-value out of it. They simply do not care what the assumptions and internal moving parts of a t-test or a linear mixed model are. I know lots of users of lmer who are focused on one and only one thing: is my effect significant? I have repeatedly seen experienced experimenters in linguistics simply ignoring the independence assumption of data points when doing a paired t-test; people often do paired t-tests on unaggregated data, with multiple rows of data points for each subject (for example). This leads to spurious significance effects, which they happily and unquestioningly accept because that was the whole goal of the exercise. I show some examples in my lecture2 slides (slide 70).
It's not just linguists, you can see the consequences of ignoring the independence assumption in
this reanalysis of the infamous study on how future tense marking in language supposedly influences economic decisions. Once the dependencies between languages are taken into account, the conclusion that Chen originally drew doesn't really hold up much:
"
When applying the strictest tests for relatedness, and when data is not
aggregated across individuals, the correlation is not significant."
Similarly,
Amy Cuddy et al's study on how power posing increases testosterone levels also got published only because the p value just scraped in below 0.05 at 0.045 or so. You can see in their figure 3 reporting the testosterone increase
that their confidence intervals are huge (this is probably why they report standard errors, it wouldn't look so impressive if they had reported CIs). All they needed to show to make their point was to get the p-value below 0.05. The practical relevance of a 12 picogram/ml increase in testosterone is left unaddressed. Another recent example from Psychological Science, which seems to publish studies that might attract attention in the popular press, is this study on
how ovulating women wear red. This study is a follow up on the
notorious Psychological Science study by
Beall and Tracy. In my opinion, the Beall and Tracy study reports a bogus result because they claim that women wear red or pink when ovulating, but when I reanalyzed their data I found that the effect was driven by pink alone. Here is my GLM fit for red or pink, red only and pink only. You can see that the "statistically significant" effect is driven entirely by pink, making the title of their paper (
Women Are More Likely to Wear Red or Pink at Peak Fertility) true only if you allow the exclusive-or reading of the disjunction:
The new study by
Eisenbruch et al reports a statistically significant effect on this red-pink issue, but now it's only about red:
"A mixed regression model confirmed that, within subjects,
the odds of wearing red were higher during the estimated
fertile window than on other cycle days, b = 0.93, p = .040,
odds ratio (OR) = 2.53, 95% confidence interval (CI) =
[1.04, 6.14]. The 2.53 odds ratio indicates that the odds of
wearing a red top were about 2.5 times higher inside the
fertile window, but there was a wide confidence interval."
To their credit, they note that their confidence interval is huge, and essentially includes 1. But since the p-value is below 0.05 this result is considered evidence for the "red hypothesis". It may well be that women who are ovulating wear red; I have no idea and have no stake in the issue. Certainly, I am not about to start looking at women wearing red as potential sexual partners (quite independent from the fact that my wife would probably kill me if I did). But it would be nice if people would try to do high powered studies, and report a replication in the same study they publish. Luckily nobody will die if these studies report mistaken results, but the same mistakes are happening in medicine, where people will die as a result of incorrect conclusions being drawn.
All these examples show why the focus on p-values is so damaging for answering research questions.
Not surprisingly, for the statistician, the main point of fitting a model (even in a confirmatory factorial analysis) is not to derive a p-value from it; in fact, for many statisticians the p-value may not even rise to consciousness. The main point of fitting a model is to define a process which describes, in the most economical way possible, how the data were generated. If the data don't allow you to estimate some of the parameters, then, for a statistician it is completely reasonable to back off to defining a simpler generative process.
This is what Gelman and Hill also explain in their 2007 book (italics mine). Note that they are talking about fitting Bayesian linear mixed models (in which parameters like correlations can be backed off to 0 by using appropriate priors; see the Stan code using lkj priors
here), not frequentist models like lmer. Also, Gelman would never, ever compute a p-value.
Gelman and Hill 2007, p. 549:
"Don’t get hung up on whether a coefficient “should” vary by group. Just allow it
to vary in the model, and then, if the estimated scale of variation is small (as with
the varying slopes for the radon model in Section 13.1), maybe you can ignore it if
that would be more convenient.
Practical concerns sometimes limit the feasible complexity of a model—for example, we might fit a varying-intercept model first, then allow slopes to vary, then add
group-level predictors, and so forth. Generally, however, it is only the difficulties of
fitting and, especially, understanding the models that keeps us from adding even
more complexity, more varying coefficients, and more interactions."
For the statistician, simplicity of expression and understandability of the model (in the Gelman and Hill sense of being able to derive sensible posterior (predictive) distributions) are of paramount importance. For the psychologist and linguist (and other areas), what matters is whether the result is statistically significant. The more vigorously you can reject the null, the more excited you get, and the language provided for this ("highly significant") also gives the illusion that we have found out something important (=significant).
This seems to be a fundamental disconnect between statisticians, and end-users who just want their p-value. A further source of the disconnect is that linguists and psychologists etc. look for cookbook methods, what a statistician I know once derisively called a "one and done" approach. This leads to blind data fitting: load data, run single line of code, publish result. No question ever arises about whether the model even makes sense. In a way this is understandable; it would be great if there was a one-shot solution to fitting, e.g., linear mixed models. It would simplify life so much, and one wouldn't need to spend years studying statistics before one can do science. However, the same scientists who balk at studying statistics will willingly spend time studying their field of expertise. No mainstream (by which I mean Chomskyan) syntactician is going to ever use commercial software to print out his syntactic derivation without knowing anything about the syntactic theory. Yet this is exactly what these same people expect from statistical software, to get an answer without having any understanding of the underlying statistical machinery.
The bottom line that I tried to convey in my course was: forget about the p-value (except to soothe the reviewer and editor and to build your career), focus on doing high powered studies, check model assumptions, fit appropriate models, replicate your findings, and publish against your own pet theories. Understanding what all these words means requires some study, and we should not shy away from making that effort.
PS I am open to being corrected---like everyone else, I am prone to making mistakes. Please post corrections, but with evidence, in the comments section. I moderate the comments because some people post spam there, but I will allow all non-spam comments.
PPS The teaching evaluation for this course just came in from ESSLLI; here it is. I believe 5.0 is a perfect score.
Statistical methods for linguistic research: Foundational Ideas (Vasishth)
| Lecturer1 | 4.9 |
|
|
| Did the course content correspond to what was proposed? | 4.9 |
| Course notes | 4.6 |
| Session attendance | 4.4 |
(19 respondents)
- Very good course.
- The lecturer was simply great. He has made hard concepts really easy to understand. He also has been able to keep the class interested. A real pity to miss the last lecture !!
- The only reason that this wasn't the best statistics course is that I had a great lecturer at my university on this. Very entertaining, informative, and correct lecture, I can't think of anything the lecturer could do better.
- Informative, deep and witty. Simply awesome.
- Professor Shravan Vasishth was hands down the best lecturer at ESSLLI 2015. I envy the people who actually get to learn from him for a whole semester instead of just a week or two. The course was challenging for someone with not much background in statistics, but Professor Vasishth provided a bunch of additional material. He's the best!
- Great course, very detailed explanations and many visual examples of some statistical phenomena. However, it would be better to include more information on regression models, especially with effects (model quality evaluation, etc) and more examples of researches from linguistic field.
- It was an extremely useful course presented by one of the best lecturers I've ever met. Thank you!
- Amazing course. Who would have thought that statistics could be so interesting and engaging? Kudos to lecturer Shravan Vasishth who managed to condense so much information into only 5 sessions, who managed to filter out only the most relevant things that will be applicable and indeed used by everyone who attendet the course and who managed to show the usefulness of the material. A great lecturer who never went on until everything was cleared up and made even the most daunting of statistical concepts seem surmountable. The only thing I'm sorry for is not having the opportunity to take his regular, semester-long statistics course so I can enjoy a more in depth look at the material and let everything settle properly. Five stars, would take again.
- Absolutely great!!